Single-cell -omics
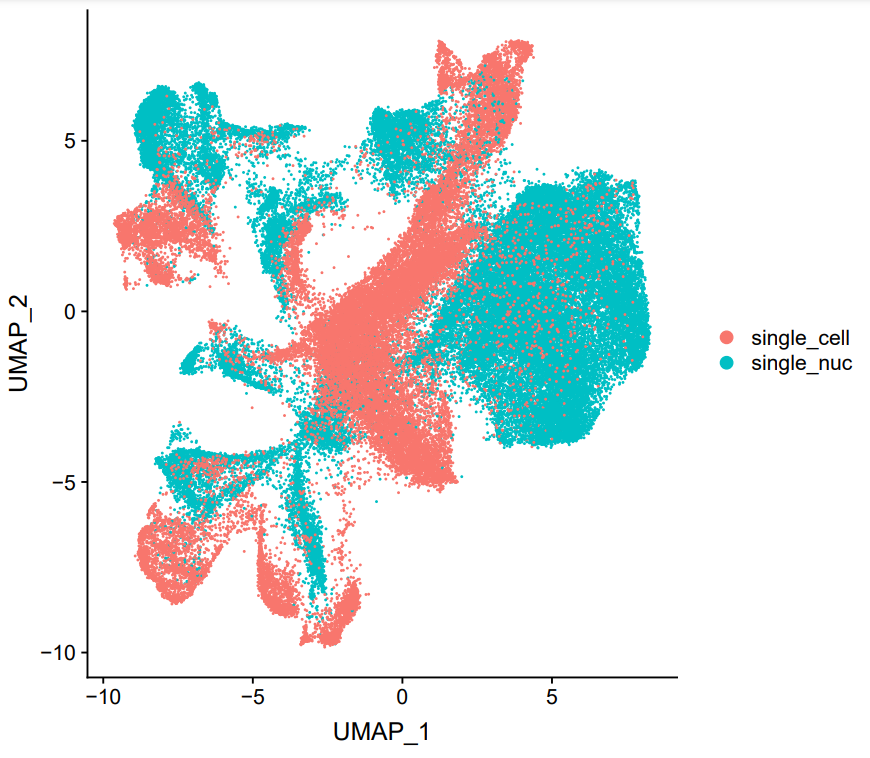
SC vs SN
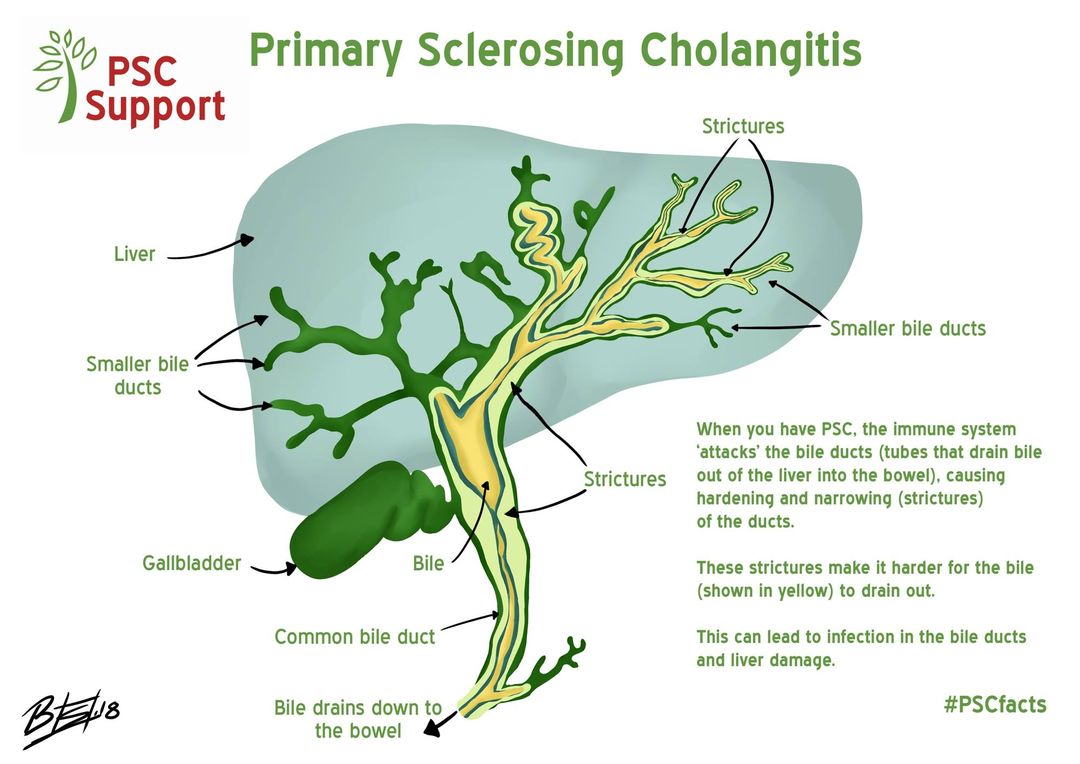
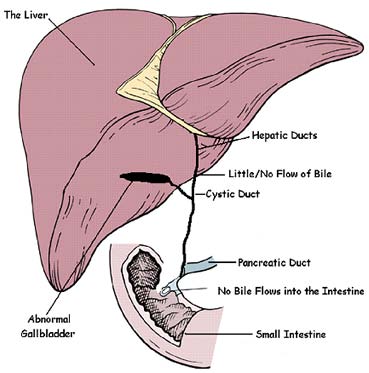
Single cell RNA sequencing has revolutionized biomedical research, however it can only be applied to fresh tissues that can be effectively dissociated into intact single cells. Frozen material such as biobanked disease cohorts is cannot be dissociated, and many diseases result in highly fibrotic or stressed cells that do not survive dissociation. To rectify this issue, many disease samples are processed using single nucleus RNA sequencing which extracts nuclei directly from tissue without requiring dissociation. However, many systematic and cell-type specific differences between single-cell and single-nucleus RNA sequencing have been observed in individual tissues, hampering efforts to compare the two. What are the differences between the RNA captured by these technologies? How can we computationally correct for them? Can we learn about post-transcriptional regulation by examining cell-type specific differences?
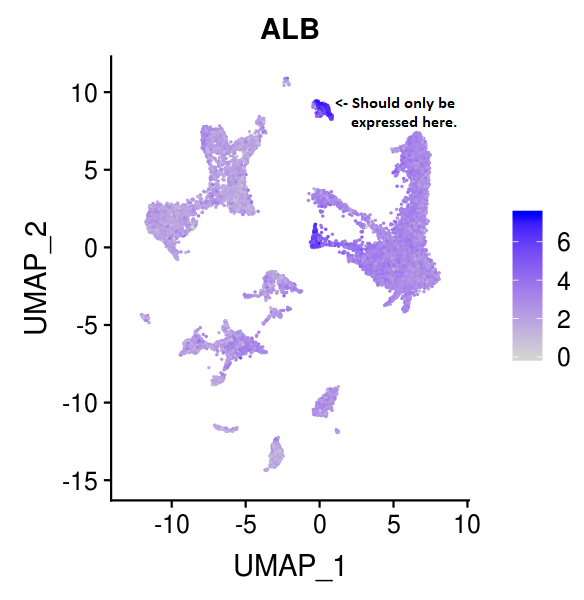
Ambient RNA Removal
During the process of tissue dissociation many cells are damaged or destroyed releasing their RNA content into the buffer that surrounds the surviving single cells. This RNA is known as 'ambient RNA' and contaminates all cells of the sample with RNA that does not belong to the captured cell. Variabililty in this contamination confounds comparisons of diseased and normal samples leading to false-positive results. Ambient RNA is a main contributor to between sample variability which causes problems when combining samples and reduces statistical power to detect differences. Ambient RNA may also cause errors in somatic variant calling when inferring tumour evolution. How do we estimate the effect of this contamination? Which tools are effective in removing it? How can we improve ambient RNA removal?
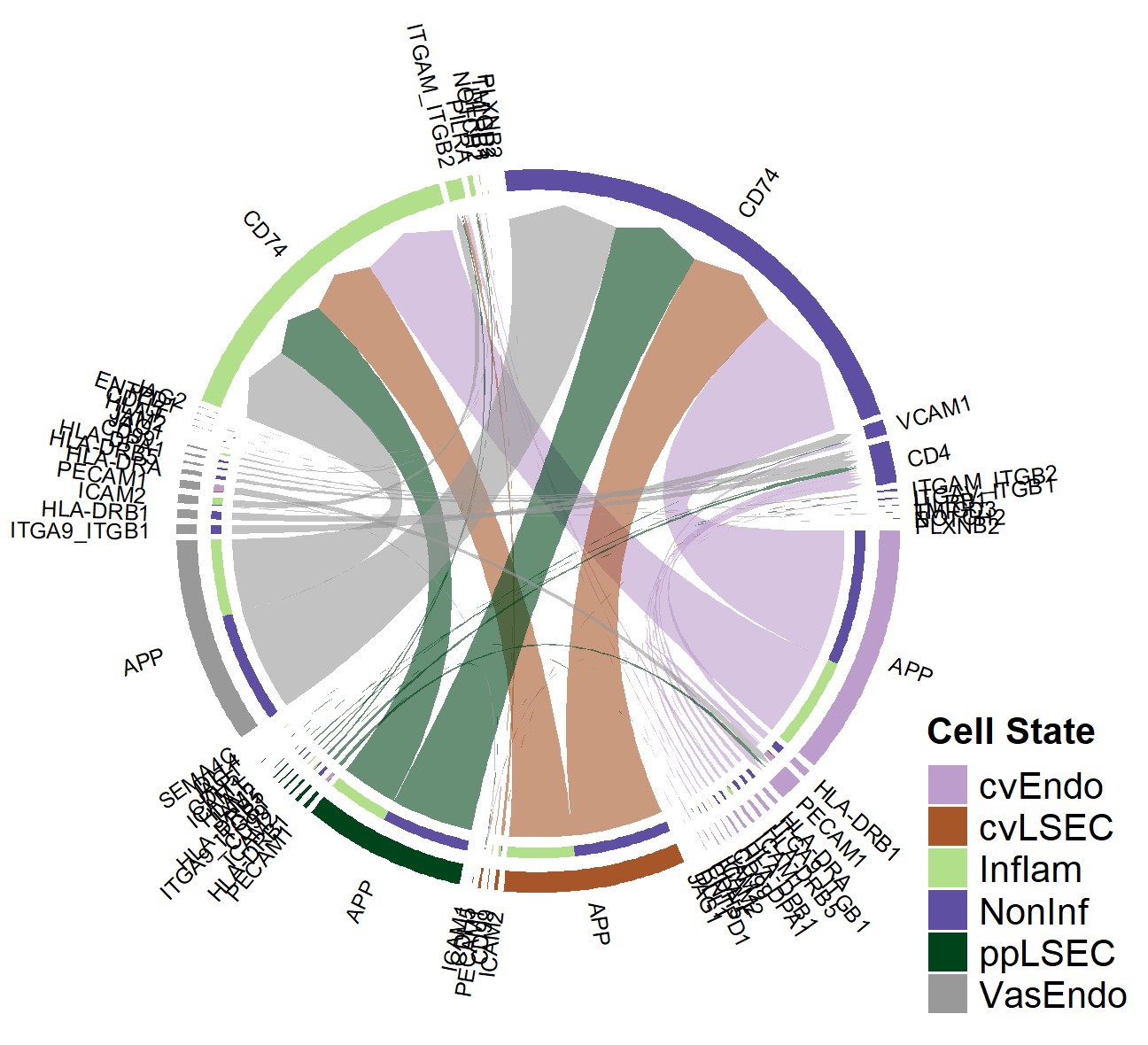
Inferring cell-cell interactions
Cells are constantly sensing and responding to their extracellular environment and nearby cells. This 'tissue microenvironment' has proven critical to the survival and proliferation of many tumours, and the progression vs resolution of inflammation-associated diseases. Recent developments in single-cell RNA sequencing enable the full characterisation tissue microenvironment in healthy and diseased tissue. However, unravelling the specific protein, signalling and metabolic interactions that regulate and maintain tissue-specific microenvironments has proven challenging due to the sparsity of single-cell RNA sequencing detection rates and paucity of characterized signalling pathways. How can we use high-throughput data to infer signalling interactions? How do we integrate single-cell RNA sequencing data with high-resolution proteomic or metabolomic data?