Research Interests and Projects
Environmental exposures, DNA methylation and breast cancer
Models of metastasis
The role of ischemia in tumor progression: implications for response to therapy.
E3: Epigenetics, embryogenesis and the environment
Research Interests and Projects
Environmental exposures, DNA methylation and breast cancer
Models of metastasis
The role of ischemia in tumor progression: implications for response to therapy.
E3: Epigenetics, embryogenesis and the environment

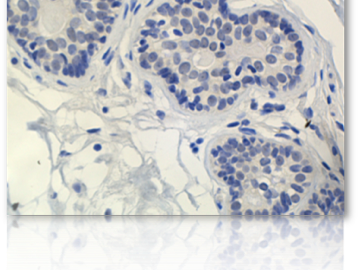
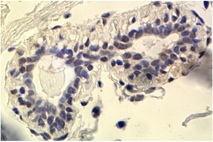

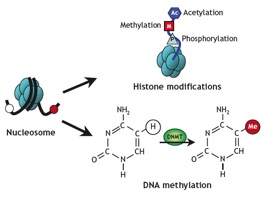
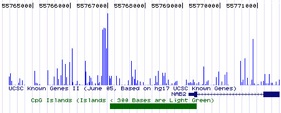



Research Summary: My research focuses on the epigenetic regulation of gene expression during tumourigenesis and development. Errors in DNA methylation patterns and histone modifications can alter gene expression, cause aberrant developmental programs and lead to inappropriate gene expression. Ongoing research in my lab focuses on (a) methylation profiling in a variety of cancer-related genes, (b) defining the functional consequences of DNA methylation changes in breast tumourigenesis in general and metastasis in particular, (c) determining how environmental carcinogens alter epigenetic patterns and contribute to tumour formation and (d) defining how environmental pollutants alter epigenetic patterns and compromise embryonic development. I am a founding member of EpiGenWestern, a collaborative research group of UWO epigenetics researchers. We share technical and scientific expertise in the areas of epigenetics, human molecular genetics, developmental biology, and cancer research.
1. Environment, DNA methylation and breast cancer. ...D. Rodenhiser (PI)
I hypothesize that environmental exposures cause changes in DNA methylation that contribute to breast tumourigenesis. Aberrant DNA methylation occurs in some tumours and cancer cell lines, but the link between environmental carcinogens, DNA methylation and breast cancer is incomplete. Environmental agents (including polycyclic hydrocarbons such as benzopyrene (B[a]P)), and micronutrients including folate, have been implicated in DNA repair and methylation events that can initiate and promoter human tumour growth. My lab is evaluating how B(a)P exposure modulates DNA methylation in breast cancer cell lines. We are (1) characterizing genome wide changes in methylation patterns (the 'methylome'), (2) identifying candidate genes with altered promoter methylation profiles and (3), assessing cellular responses and molecular mechanisms to B(a)P by defining its effects on the expression of BRCA1 and the DNA methyltransferase genes (DNMT1, 3a and 3b).
2. Models of Metastasis ... Check out our website: the metastasisproject.ca
Metastasis is the main cause of death in breast cancer. Although early detection and management of breast cancer in general has improved, it continues to be a challenge to determine which patients are at highest risk of metastasis, to obtain early clinical detection of metastases, and to manage these patients appropriately. Advances in gene profiling and proteomics have yielded "signature" patterns that appear to be associated with invasiveness, metastasis in general and site-specific metastasis. Model systems are thus needed to assess metastasis-related genes as targets for new therapeutic approaches. My role in this project is to identify and characterize epigenetic changes in target genes relevant to breast cancer progression and metastasis. We will develop a precise map of promoter hypermethylation and transcriptional inactivation of these genes in the context of progression. We will also assess promoter methylation profiles in the context of survival/growth/invasion of these cells in the lymph node environment. Once epigenetically genes are identified, the effects of agents affecting methylation status on gene expression in the lymph node environment will be tested.
3. The role of ischemia in tumor progression: implications for response to therapy.
BL Coomber, Univ Guelph (PI); D. Rodenhiser (Co-applicant)
We are exploring how anti-angiogenic cancer therapy (which prevents cancer growth and spread by preventing
blood vessels from growing into the tumor) might change cancer cells in unexpected ways. We are focusing on
the fact that tumors develop regions of low oxygen (hypoxia), a situation known to lead to DNA mutations, and
changes in gene production. Since anti-angiogenic therapy works by preventing new blood vessels from
growing into the tumor, it probably also increases the amount of hypoxia in the tumors, and this may lead to
unexpected changes in the treated cancers.
4. E3: Epigenetics, environment and embryogenesis ...D. Rodenhiser (PI)
Embryonic development is a carefully choreographed process during which genes are turned on (and off), organs grow, limbs are formed and eventually an offspring is born. Unfortunately, normal development is constantly challenged by (1) chemicals present in the environment, (2) naturally produced chemical breakdown products and (3), fluctuations in the diet. As a consequence, developmental programming can go wrong, leading to decreases in fertility, abnormal offspring, miscarriage or even harmful effects only seen in later generations. This study, submitted to NSERC, will characterize changes in developing embryos caused by chemicals found in pollution. I will use mouse embryos to identify and characterize the interactions between the environment and DNA methylation patterns during development. My research program will use molecular technologies to (1) detect epigenetic and gene expression changes in embryos following chemical exposures, (2) identify genes that can be turned on/off by epigenetic alterations and (3), define methylation biomarkers of toxicity that could be used in environmental safety assessment. Practical applications of this research include managing reproductive health in animals, improving livestock production and yield through optimal nutrition as well as maintaining the diversity and sustainability of wildlife populations in natural ecosystems. This research could eventually lead to bioassay tests to allow toxicity testing or chemical risk assessment in human, livestock and wildlife.
Info for
Prospective
Students: